There’s actually two statements that I’d like to deal with here. The first is, of course, that most mutations are harmful. The second is that selection only removes alleles from the gene pool and therefore removes the variation that exists because of mutations. These two together or either separately is used as an attack on evolution. Both of these charges are wrong and the topics are closely related so here we go.
Mutations, as we all know (and I’ve only met a very few people who don’t think mutations even happen), are changes in the DNA of an organism. It can be something as minor as a single nucleotide in the DNA changing or as major as the loss or gain of an entire chromosome. The average rate for mutations in humans is something on the order of 175 mutations per diploid genome per generation. This link also tells us that substitution mutations are 4 times as likely to occur as deletions or insertions.
A substitution is when one nucleotide (A,C,G, or T) is replaced by another letter. A deletion removes a nucleotide completely from a gene, while an insertion puts one in. Since most mutations are substitutions, it can easily be shown that the vast majority of mutations do not even change the resulting protein. The reason is shown below.
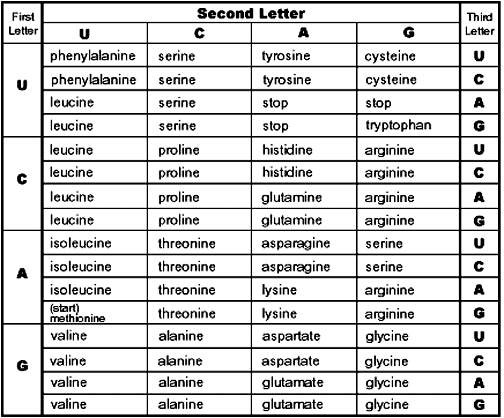
This is a codon chart. DNA is transcribed into messenger RNA (mRNA) that can leave the nucleus. A ribosome then translates the mRNA into a protein. THe ribosome reads three nucleotides at a time. This three nucleotide set is called a codon. Each codon has a unique amino acid that matches it. (The process is more complicated than this, but this will get us to where we need to be today.)
There are four nucleotides in RNA (A, C, G, and U). There are three spots in the codon. Some simple math tells us that there are 64 possible combinations of nulceotides. But there are only 20 amino acids in use in the human body. There’s also a STOP codon that tells the ribosome to stop adding amino acids. So, we only need 21 spots, but we have 64. The result is that many amino acids have multiple codons that will code for them.
For example, from the chart, we see that leucine (something of an overachiever) has six codons that will work for it: UUA, UUG, UCU, UCA, UCG, and UCC.
Now, that means that the particular codon can be hit with a variety of substitution mutations and it will not change the amino acid that fits in that spot. No change at all. Think of this like using a thesaurus. To describe Rocket Raccoon one might use the word short, tiny, diminutive, petite, runt, or any of a dozen other words that mean pretty much the same thing. A writer might choose a particular word for a reason, but any word will describe the height (or lack thereof) of the character.
This gets even more interesting.
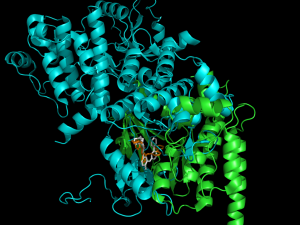
This is a protein (actually, it’s two proteins). Click to enlarge the picture. You see that little area of orange and white in the lower center? That is what is called the ‘active site’. That’s the part of the protein that does all the work. It may store an oxygen molecule, it may change ATP into ADP, it may bind two amino acids together, or any other of a million possible things. But that miniscule area is what is critical to the function of the protein. I’ll note here that proteins may have more than one function.
But anyway, every part of the protein that is not the active site only has one function. To make the protein fold into that shape. That’s all. A lot of the amino acids come in families with similar chemical or physical properties. Some (three) have a positive charge in a particular spot. Some (two) have a negative charge in that same spot. Some are attracted to water and some are repelled by water. In quite a few cases, it doesn’t matter what amino acid is present… provided that it meets one of those criteria.
For example, arginine, histidine, and lysine all have a positive side chain. If it’s not in the active site, then it may be likely that any of those three will do in that spot. Now, there are ten possible codons that will produce the same result. Any mutation that changes from histidine to arginine won’t change the protein in any appreciable way.
I will say that it is possible for these kinds of mutations to set-up larger changes in the future. These are called potentiating mutations. For example, in Lenski’s work, a certain mutation happened long before the E. coli developed the ability to utilize citrate. If that mutation doesn’t happen first, then the remaining changes still don’t allow citrate forms to evolve.
It gets even easier, because, even if the active site changes, the protein may still work. There can be different versions of a gen called an allele. These alleles may do something similar, but not exactly the same. One allele might give you a widow’s peak, while the other gives you a straight hairline.
There are several with many possible alleles. Due to genetics, you only have two of them (one from mom and one from dad… unless one of them mutated, then you may be unique). The HLA-B gene is also something of an overachiever. There are 3,285 alleles that construct 2,459 unique proteins. All of these proteins re similar enough to do the same thing (help the immune system recognize the difference between your cells and invading cells). You could have dozens of mutations in this gene and the protein would be perfectly fine. It’s very amenable to change… some proteins aren’t so amenable to change.
Don’t get me wrong. It’s entirely possible that a mutation occurs which kills the organism before it’s first cell division. That happens. Indeed, it probably happens much more frequently than anyone ever realizes (it takes weeks before a human female even knows she’s pregnant).
But mutations aren’t the massively detrimental problem that many people seem to think.
Which brings us to part two of our story. Selection is a removing factor. That’s true.
But when you have 3,285 alleles to choose from and, on average, new ones are being created every day. If one is detrimental, then it’s gone. But so many more have no effect at all (coding for the exact same amino acid) or an effect which is effectively invisible (coding for the same type of amino acid), that there are way more potential alleles in a population than selection can remove.
In other words, mutation adds more alleles to the population faster than selection can remove them. This is because many of the changes to the genes are simply too minuscule to have a selection effect.
This paper[1] describes it very well. The authors show that, given certain circumstances, that a population will tend to express multiple phenotypes. This isn’t what Darwin predicted, the old ‘survival of the fittest’, but it is, instead, ‘survival of the good enough’. In the real world, there may be multiple ways to survive. In a gross sense, think of one species dealing with a competitor by becoming smaller, faster, and hiding more and another species becoming larger, stronger, and tougher. This paper is talking about alleles, not large scale features like that, but it’s an analogy to help you understand what I’m describing.
Both solutions may be equally effective. And some of the population will go one way, some the other way, based on what is already present in their genome from all those mutations that have come before.
I have little hope that this will put to bed these creationist tropes. But still, it’s fascinating to go beyond high school biology, where you might have two traits, each with two options. In the real world, we have thousands of traits, some with thousands of options and mutations which generate more and more options as time goes on.
__________________________________
[1] Schaper, S., Louis, A. A. & Rutherford, S. The Arrival of the Frequent: How Bias in Genotype-Phenotype Maps Can Steer Populations to Local Optima. PLoS ONE 9, (2014).